-
Hemoglobin switch
-
TF kinetics
-
Enhancers and condensates
How does BCL11A regulate hemoglobin switching?
Human hemoglobin genes undergo developmental switching during ontogeny. The fetal-to-adult switch occurs a few months after birth. This process transitions hemoglobin from the high oxygen-affinity fetal type to the more regulatable adult type. Human α- and β-globin loci have been paradigms for studying spatiotemporal gene regulation for decades. Understanding how this switching occurs not only deepens our understanding of gene regulation but also holds great clinical relevance. Hemoglobinopathies, including β-thalassemia and sickle cell disease, are caused by mutations in the adult hemoglobin gene. Reversal of hemoglobin switching and reactivation of fetal hemoglobin have been proven effective in treating β-thalassemia and sickle cell disease.
Transcription factor BCL11A was identified as a major repressor of fetal γ-globin in 2008 through GWAS. With the development of CRISPR/Cas9 technology, researchers have investigated targeting BCL11A using CRISPR. This has led to the first gene-editing therapy, which uses targeted genome editing of human somatic cells to cure disease. This historic therapy, casgevy, was approved in 2023.
Our previous work focused on the mechanistic aspects of BCL11A action, specifically how BCL11A represses fetal globin genes. Our main discoveries are as follows:
- BCL11A binds to a distal TGACCA motif in the γ-globin promoters to repress its expression. Interestingly, human genetic variants located in or overlapping with this motif have been known to cause fetal globin reactivation for over 30 years, but the mechanism remained a mystery. This work solved this human genetic puzzle.
- BCL11A competes with transcription activators, including NF-Y, to repress γ-globin. We have proposed a transcription factor competition model, suggesting that competition between a stage-selective repressor (BCL11A) and a ubiquitous activating factor (NF-Y) within a discrete 35 bp region that encompasses duplicated TGACCAAT boxes directs the developmental sequence of globin gene expression across the entire β-globin locus.
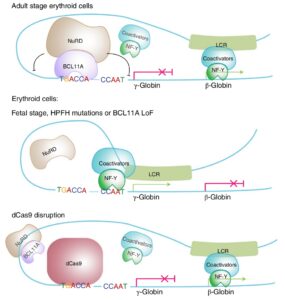
Liu et al, Cell 2018; Liu et al, Nature Genetics 2021
- Using base editing, we modified the TGACCA motif and reactivated γ-globin genes. This serves as a proof-of-principle for base editing as a therapeutic strategy. Biotechs exploring this strategy by targeting this motif worldwide have demonstrated its efficacy. Tens of patients have been cured in clinical trials by editing this TGACCA motif.
- We were early adopters of the CUT&RUN methodology, which has played a pivotal role in profiling BCL11A chromatin occupancy. We collaborate with G-C Yuan's lab and developed a widely used software package, CUTRUNTools, for convenient analysis of CUT&RUN data, including mapping, peak calling, footprint analysis, etc.
- We optimized the CUT&RUN library preparation procedure. The protocol has been deposited on protocols.io and has been widely used (>50,000 views).
How do the kinetic behaviors of transcription factors determine their function?
Transcription factors (TFs) are a class of molecules that regulate gene expression by directly recognizing and binding to specific sequence motifs within regulatory regions. The binding of TFs to their cognate DNA motifs is a thermodynamic process. Under equilibrium conditions, this process involves frequent binding and dissociation events, primarily dictated by the affinity of the TF for DNA. Consequently, the fraction of TF molecules is bound to chromatin, and the duration of chromatin residence vary among different factors. Additionally, the speed at which a TF freely diffuses within the nucleoplasm is determined by its physical properties. These factors collectively define the kinetic behaviors of TFs, including bound fractions, chromatin residence time, and diffusion coefficients, etc. These fundamental physical properties influence how rapidly a TF locates its target and how efficiently it executes its function.
Counterintuitively, TFs exhibit greater dynamism than one might expect. Seminal studies in the early 2000s demonstrated that most TFs have low bound fractions and strikingly short chromatin residence times (2–30 seconds). This highly dynamic behavior has been partially attributed to the action of chromatin remodelers and proteasomes, which function as kinetic proofreading mechanisms to overcome the Hopfield barrier — a thermodynamic constraint on information processing that requires high precision. These characteristics are closely related to TF target search and transcriptional bursting.
Using hemoglobin switching as a model system (as well as other available models), we delve into the kinetic behavior of TFs and TF-DNA interactions to understand how these behaviors determine transcription.
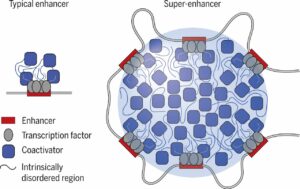
What makes an enhancer condensate?
Enhancers are cis-regulatory elements (CREs) that control gene expression from a distance. It is estimated that there are > 1 million putative enhancers in the human genome, a number that far exceeds the total number of genes. A significant proportion of pathogenic human genetic variants are located within enhancers.
Last modified: May 2025